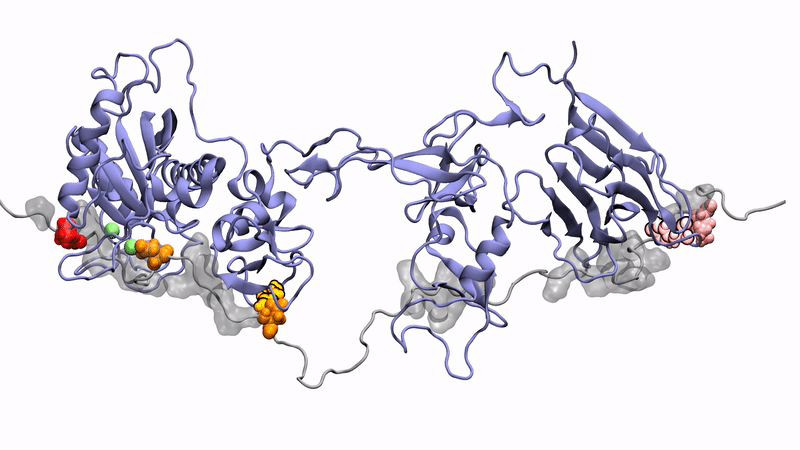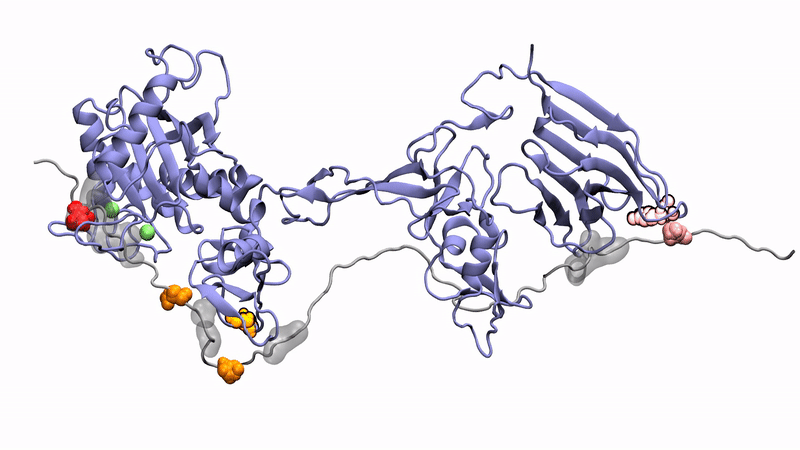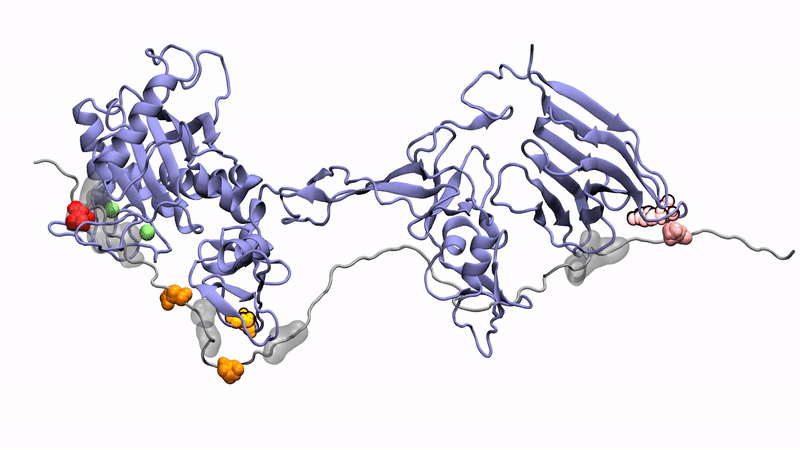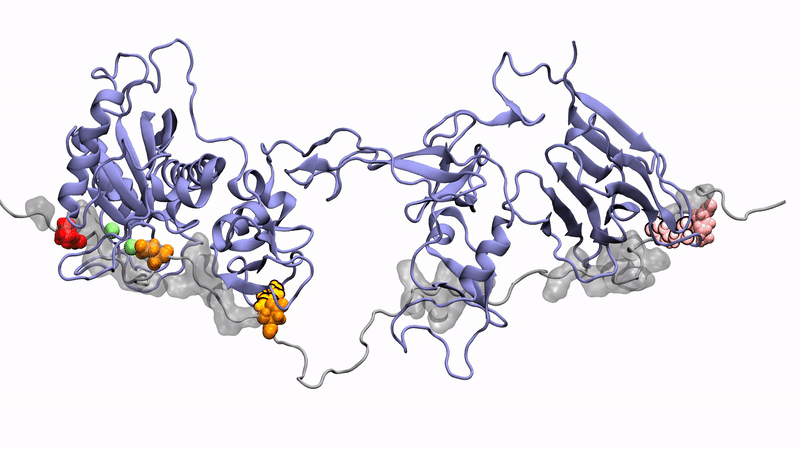
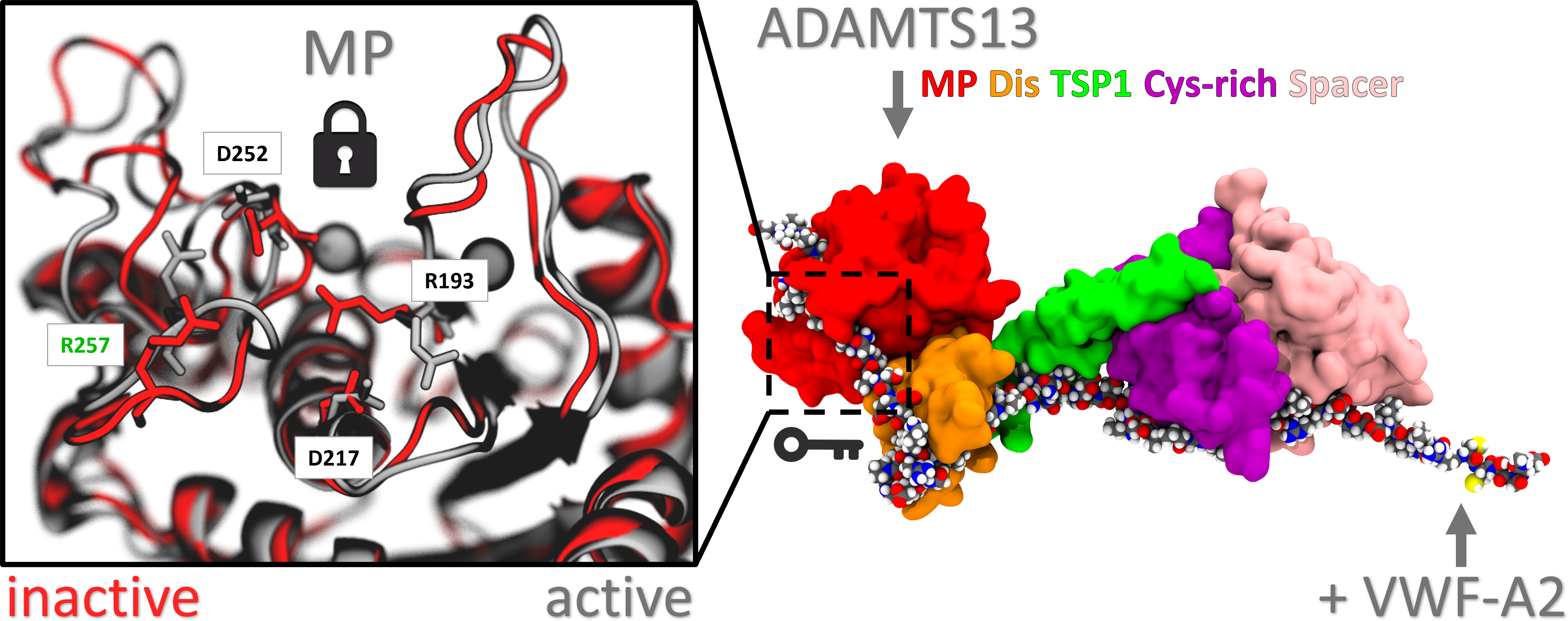
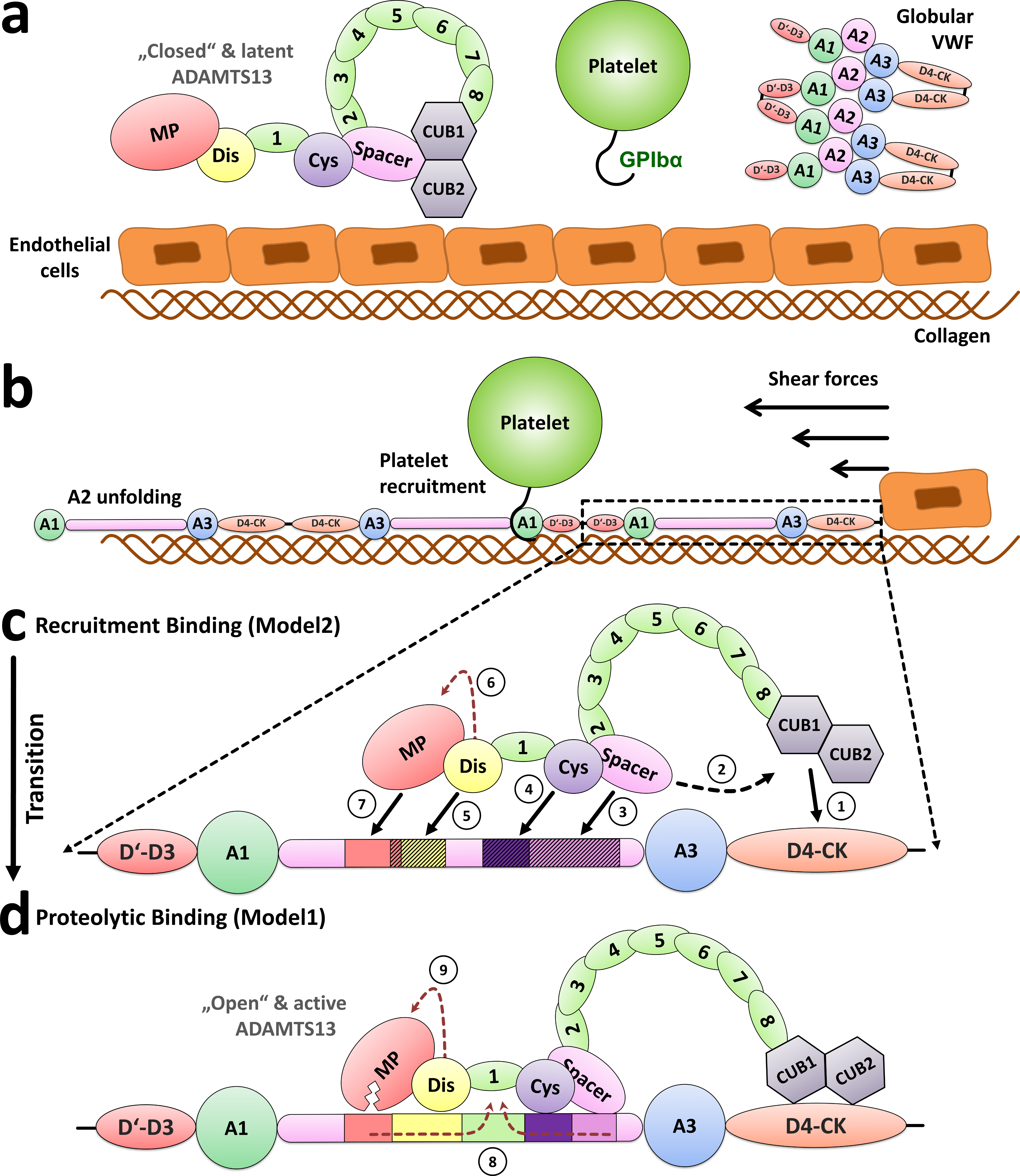
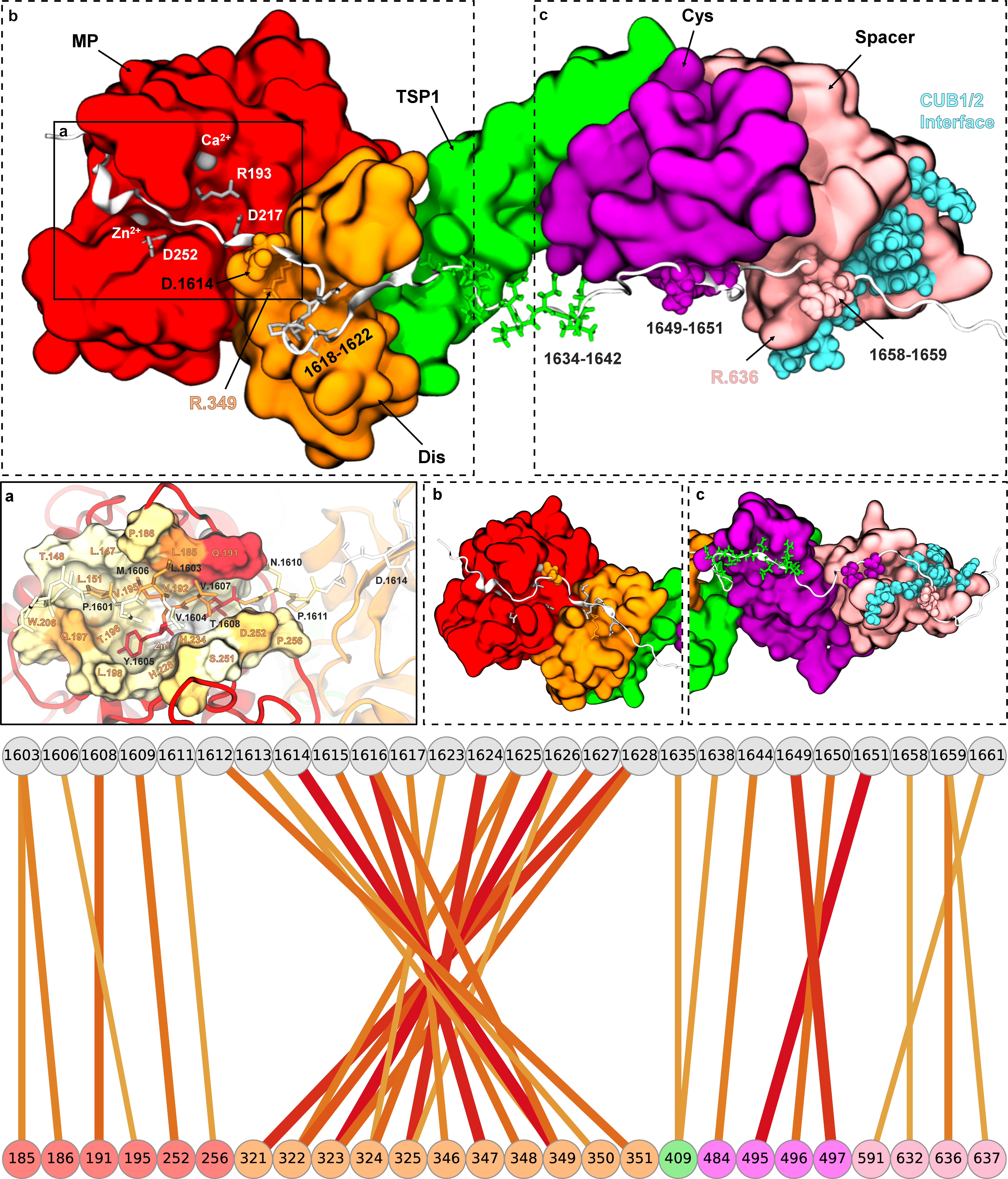
Norman's paper named "Molecular interplay of ADAMTS13-MDTCS and von willebrand Factor-A2: deepened insights from extensive atomistic simulations" is published in the Journal of Biomolecular Structure and Dynamics. Congratulations Norman!
Supporting data sets on Zenodo and code repository for TIGR2hs (PE) on Github https://github.com/SLx64/TIGER2hs